
Ordinary Least Squares & the Normal Equations¶
The Closed-Form Shortcut to the Best-Fit Line¶
Gradient descent finds the minimum by walking downhill one step at a time. But for linear regression there is a one-shot algebraic solution that jumps directly to the bottom: the Normal Equations. This notebook derives it, implements it, and explains when to use it versus gradient descent.
From Gradient to Zero¶
In the gradients notebook we established the matrix gradient of the MSE loss:
Gradient descent keeps subtracting this until it is small. But what if we set it exactly to zero and solve?
If is invertible, multiply both sides by its inverse:
This is the Normal Equation — the exact minimiser of MSE, computed in one step.
Visual: Two Paths to the Same Minimum¶
Matrix Dimensions Reference¶
For samples and features (plus a bias column):
| Symbol | Meaning | Shape |
|---|---|---|
| Design matrix (with bias column of 1s) | ||
| Target vector | ||
| Gram matrix | ||
| Inverse of Gram matrix | ||
| Moment vector | ||
| Optimal coefficients (incl. intercept) |
OLS from Scratch — NumPy Implementation¶
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# Toy advertising dataset
tv_spend = np.array([230, 44, 17, 151, 180, 8, 57, 120, 200, 100])
sales = np.array([22, 10, 7, 18, 20, 3, 11, 15, 21, 13])
# Build design matrix X with bias column
n = len(tv_spend)
X = np.column_stack([np.ones(n), tv_spend]) # shape (n, 2)
# Normal Equations: theta* = (X^T X)^{-1} X^T y
theta = np.linalg.inv(X.T @ X) @ X.T @ sales
intercept, slope = theta
print(f"Intercept (theta_0): {intercept:.4f}")
print(f"Slope (theta_1): {slope:.4f}")
print(f"Interpretation: baseline sales = {intercept:.2f}; each $1 in TV adds {slope:.4f} in sales")
# Plot
x_line = np.linspace(0, 250, 200)
fig, ax = plt.subplots(figsize=(7, 4))
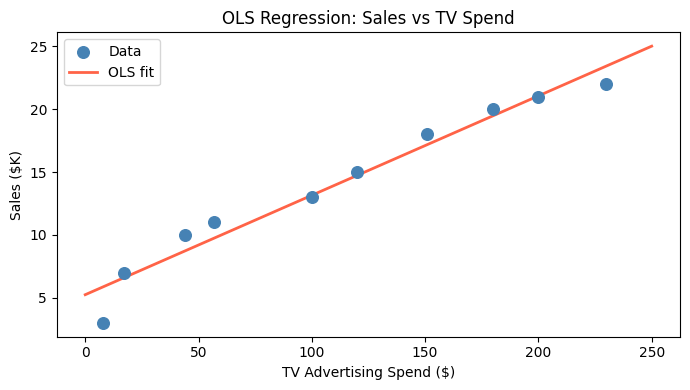
ax.scatter(tv_spend, sales, color='steelblue', s=70, zorder=3, label='Data')
ax.plot(x_line, intercept + slope * x_line, color='tomato', linewidth=2, label='OLS fit')
ax.set_xlabel('TV Advertising Spend ($)')
ax.set_ylabel('Sales ($K)')
ax.set_title('OLS Regression: Sales vs TV Spend')
ax.legend()
plt.tight_layout()
plt.show()Intercept (theta_0): 5.2428
Slope (theta_1): 0.0791
Interpretation: baseline sales = 5.24; each $1 in TV adds 0.0791 in sales
OLS with scikit-learn¶
For production use, LinearRegression calls LAPACK’s least-squares solver internally.
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score, mean_squared_error
rng = np.random.default_rng(42)
n = 80
X_feat = rng.uniform(0, 10, (n, 2))
true_coef = np.array([1.5, -0.8])
y = X_feat @ true_coef + 3.0 + rng.normal(0, 1.2, n)
model = LinearRegression().fit(X_feat, y)
y_hat = model.predict(X_feat)
print(f"Intercept: {model.intercept_:.4f} (true: 3.0)")
print(f"Coef[0]: {model.coef_[0]:.4f} (true: 1.5)")
print(f"Coef[1]: {model.coef_[1]:.4f} (true: -0.8)")
print(f"R²: {r2_score(y, y_hat):.4f}")
print(f"RMSE: {np.sqrt(mean_squared_error(y, y_hat)):.4f}")
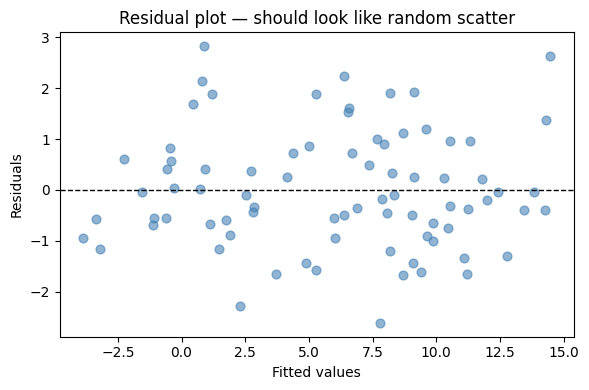
# Residual plot
fig, ax = plt.subplots(figsize=(6, 4))
ax.scatter(y_hat, y - y_hat, alpha=0.6, color='steelblue', s=40)
ax.axhline(0, color='black', linestyle='--', linewidth=1)
ax.set_xlabel('Fitted values')
ax.set_ylabel('Residuals')
ax.set_title('Residual plot — should look like random scatter')
plt.tight_layout()
plt.show()Intercept: 2.9335 (true: 3.0)
Coef[0]: 1.5645 (true: 1.5)
Coef[1]: -0.8477 (true: -0.8)
R²: 0.9458
RMSE: 1.1508
Full Diagnostic Summary with statsmodels¶
scikit-learn optimises for prediction. For inference — p-values, confidence intervals, VIF, autocorrelation tests — use statsmodels.
The cell below contains a reusable regression_summary_paragraph helper that formats the key diagnostics in plain language.
import pandas as pd
import statsmodels.api as sm
from statsmodels.stats.outliers_influence import variance_inflation_factor
def regression_summary_paragraph(model,
alpha: float = 0.05,
vif_thresholds: tuple = (5, 10),
r2_thresholds: tuple = (0.3, 0.6)) -> str:
"""
Creates a readable, multi-line summary from a fitted statsmodels OLS model.
Returns a short, human-readable multi-line string (suitable for printing in
notebooks or terminals). Keeps the same numeric checks as before but places
results on separate lines for better readability.
"""
n = int(model.nobs)
r2 = model.rsquared
adj_r2 = model.rsquared_adj
fval = model.fvalue
fp = model.f_pvalue
# R² strength
if r2 < r2_thresholds[0]:
r2_qual = "weak"
elif r2 < r2_thresholds[1]:
r2_qual = "moderate"
else:
r2_qual = "strong"
overall_sig = "statistically significant" if fp < alpha else "not statistically significant"
# Coefficients (one line per predictor)
coef_lines = []
conf = model.conf_int(alpha).to_numpy()
for var, coef, pval, ci_low, ci_high in zip(
model.params.index,
model.params,
model.pvalues,
conf[:, 0],
conf[:, 1],
):
if var == "const":
continue
sig_text = "significant" if pval < alpha else "not significant"
coef_lines.append(
f"{var}: {coef:+.3f} (p={pval:.4f}, 95% CI [{ci_low:.3f}, {ci_high:.3f}]) — {sig_text}"
)
# VIF (if applicable)
vif_line = ""
if model.df_model > 0:
X = model.model.exog
if X.shape[1] > 1: # more than just intercept
vif_df = pd.DataFrame({
"variable": model.model.exog_names,
"VIF": [variance_inflation_factor(X, i) for i in range(X.shape[1])],
})
vif_df = vif_df[vif_df["variable"] != "const"]
vif_items = []
for _, row in vif_df.iterrows():
v = row["VIF"]
if v < vif_thresholds[0]:
q = "low"
elif v < vif_thresholds[1]:
q = "moderate"
else:
q = "high"
vif_items.append(f"{row['variable']} = {v:.2f} ({q})")
if vif_items:
vif_line = "VIFs: " + ", ".join(vif_items)
# Durbin-Watson
dw = sm.stats.durbin_watson(model.resid)
dw_interp = (
"approx 2 — no clear autocorrelation"
if 1.5 <= dw <= 2.5
else (f"{dw:.2f} — possible positive autocorrelation" if dw < 1.5 else f"{dw:.2f} — possible negative autocorrelation")
)
# Assemble multi-line output
lines = []
lines.append(f"Observations: {n} | R2 = {r2:.3f} (adj R2 = {adj_r2:.3f}) -- {r2_qual}")
lines.append(f"Overall model: {overall_sig} | F = {fval:.2f}, p = {fp:.4f}")
lines.append("")
lines.append("Coefficients:")
if coef_lines:
for cl in coef_lines:
lines.append(f" - {cl}")
else:
lines.append(" (no predictors besides intercept)")
if vif_line:
lines.append("")
lines.append(vif_line)
lines.append("")
lines.append(f"Durbin-Watson: {dw_interp}")
return "\n".join(lines)# --- Minimal runnable example using synthetic data
import numpy as np
import pandas as pd
np.random.seed(0)
n = 200
# create a small advertising-style dataset (predictors only)
your_predictors_df = pd.DataFrame({
"TV": np.random.normal(150, 30, size=n),
"Radio": np.random.normal(30, 8, size=n),
"Online": np.random.normal(50, 15, size=n),
})
# generate a target with known coefficients + noise
dependent_series = (
10.0
+ 0.05 * your_predictors_df["TV"]
+ 0.4 * your_predictors_df["Radio"]
- 0.02 * your_predictors_df["Online"]
+ np.random.normal(0, 3.0, size=n)
)
dependent_series = pd.Series(dependent_series, name="Sales")
# Fit OLS and show the human-readable paragraph
X = sm.add_constant(your_predictors_df)
model = sm.OLS(dependent_series, X).fit()
print("\nReadable summary:\n")
print(regression_summary_paragraph(model))
Readable summary:
Observations: 200 | R2 = 0.551 (adj R2 = 0.544) -- moderate
Overall model: statistically significant | F = 80.05, p = 0.0000
Coefficients:
- TV: +0.048 (p=0.0000, 95% CI [0.035, 0.061]) — significant
- Radio: +0.369 (p=0.0000, 95% CI [0.313, 0.424]) — significant
- Online: -0.025 (p=0.0657, 95% CI [-0.052, 0.002]) — not significant
VIFs: TV = 1.01 (low), Radio = 1.04 (low), Online = 1.04 (low)
Durbin-Watson: approx 2 — no clear autocorrelation
How to read the summary output
| Field | What it tells you |
|---|---|
| R² | Fraction of target variance explained (0–1; higher = better fit) |
| Adj R² | R² penalised for extra features — use for model comparison |
| F-stat / p | Is the model as a whole better than just predicting the mean? |
| Coef p-value | Is this predictor’s contribution distinguishable from noise? |
| 95% CI | Range where the true coefficient lies with 95% confidence |
| VIF | Variance inflation factor: VIF > 10 signals severe multicollinearity |
| Durbin-Watson | Tests residual autocorrelation; ~2 is ideal, <1.5 or >2.5 is a red flag |
In the example above, TV and Radio are significant, Online is not (p > 0.05). VIFs are all near 1, confirming no multicollinearity. Durbin-Watson near 2 means residuals are independent.
The Four OLS Assumptions¶
OLS gives the Best Linear Unbiased Estimator (BLUE) when these hold (Gauss-Markov theorem):
| Violation | Consequence | Fix |
|---|---|---|
| Non-linearity | Biased coefficients | Add polynomial terms or transform features |
| Autocorrelated residuals | Underestimated standard errors | Use time-series models (ARIMA) or GLS |
| Heteroscedasticity | Wrong confidence intervals | Use robust standard errors or WLS |
| Multicollinearity (VIF > 10) | Unstable, inflated coefficients | Drop or combine collinear features; use Ridge |
| Outliers | Distorted fit | Investigate; use robust regression or winsorise |
When to Use Normal Equations vs Gradient Descent¶
Try It in the Browser¶
Compute the Normal Equations solution from scratch using only Python built-ins.
Guided Practice¶
What is the main goal of Ordinary Least Squares?¶
Why is the Normal Equation impractical for very large feature spaces (p >> 10 000)?¶
A VIF of 14 for a predictor signals what problem?¶
A Durbin-Watson statistic of 0.8 suggests what?¶
Exercises¶
Exercise 1 — Normal Equations by hand¶
Given samples: , .
Build the design matrix (add a column of ones).
Compute and .
Solve for .
Compare with scikit-learn’s
LinearRegression.
import numpy as np
from sklearn.linear_model import LinearRegression
x = np.array([1.0, 2.0, 3.0, 4.0])
y = np.array([2.0, 4.0, 5.0, 4.0])
# TODO: build X, compute Normal Equations solution, compare to sklearn
Solution
X = np.column_stack([np.ones(len(x)), x])
theta_ne = np.linalg.inv(X.T @ X) @ X.T @ y
print("Normal Equations:", theta_ne)
lr = LinearRegression().fit(x.reshape(-1,1), y)
print("sklearn intercept:", lr.intercept_, "slope:", lr.coef_[0])Both should give intercept ≈ 1.5, slope ≈ 0.8.
Exercise 2 — Detect and interpret multicollinearity¶
Generate a dataset where two features are highly correlated (r > 0.95). Fit OLS, inspect the VIF values and coefficient confidence intervals. Describe what you observe.
import numpy as np
import pandas as pd
import statsmodels.api as sm
from statsmodels.stats.outliers_influence import variance_inflation_factor
rng = np.random.default_rng(5)
n = 100
x1 = rng.normal(0, 1, n)
x2 = x1 + rng.normal(0, 0.05, n) # x2 almost identical to x1
y = 2*x1 + 3 + rng.normal(0, 1, n)
df = pd.DataFrame({'x1': x1, 'x2': x2})
X = sm.add_constant(df)
# TODO: fit OLS, print regression_summary_paragraph, observe VIF and CI width
Exercise 3 — OLS vs Gradient Descent speed comparison¶
Generate a dataset with rows and features. Time both approaches (Normal Equations via np.linalg.lstsq and your gradient descent implementation) and confirm they converge to the same .
import numpy as np, time
rng = np.random.default_rng(99)
n, p = 500, 50
X_raw = rng.normal(0, 1, (n, p))
X = np.column_stack([np.ones(n), X_raw])
true_theta = rng.uniform(-2, 2, p+1)
y = X @ true_theta + rng.normal(0, 0.5, n)
# TODO: time Normal Equations solution
# TODO: time gradient descent (use the matrix form from gradients.ipynb)
# TODO: compare max absolute difference between the two theta vectors
/var/folders/93/7lt42x5j7m39kz7wxbcghvrm0000gn/T/ipykernel_67928/3962639806.py:8: RuntimeWarning: divide by zero encountered in matmul
y = X @ true_theta + rng.normal(0, 0.5, n)
/var/folders/93/7lt42x5j7m39kz7wxbcghvrm0000gn/T/ipykernel_67928/3962639806.py:8: RuntimeWarning: overflow encountered in matmul
y = X @ true_theta + rng.normal(0, 0.5, n)
/var/folders/93/7lt42x5j7m39kz7wxbcghvrm0000gn/T/ipykernel_67928/3962639806.py:8: RuntimeWarning: invalid value encountered in matmul
y = X @ true_theta + rng.normal(0, 0.5, n)
Common Pitfalls¶
Summary¶
Key takeaways
| Concept | One-line meaning |
|---|---|
| Normal Equations | — exact MSE minimiser |
| Derivation | Set MSE gradient to zero; solve algebraically |
| Complexity | for inversion — fast for small , slow for large |
| vs GD | No iterations, no learning rate — but not scalable to huge feature spaces |
| R² / Adj R² | Variance explained; adj R² penalises extra features |
| VIF | Multicollinearity check; VIF > 10 = problem |
| Durbin-Watson | Autocorrelation check; ~2 = ideal |
| BLUE property | OLS gives the best linear unbiased estimator when all 4 assumptions hold |
Next Up — Polynomial Features¶

OLS gives the best straight-line fit. But what if the relationship curves?¶
The next notebook — Polynomial Features — shows how to extend the linear model to capture non-linear patterns by adding $x^2$, $x^3$, and interaction terms as new features. The model stays linear in $\boldsymbol{\theta}$, so OLS still applies — but the decision boundary becomes a curve.
Dependencies you already have: the design matrix $\mathbf{X}$, coefficient interpretation, and the OLS solution formula. Polynomial features just expand $\mathbf{X}$ with new columns.